Help
Below you can find a description of how to use 2Struc and 2StrucCompare, as well as an FAQ.
Submitting a job
Input: One PDB file uploaded from your computer or one PDB 4-letter code, then click Submit.
Results Page
-
Choose which chain you wish to see the results for using the dropdown menu on the top-left.
-
Below this you can choose to include or remove missing residues from the analysis. These are automatically parsed from the PDB header if present and are classified as "other".
-
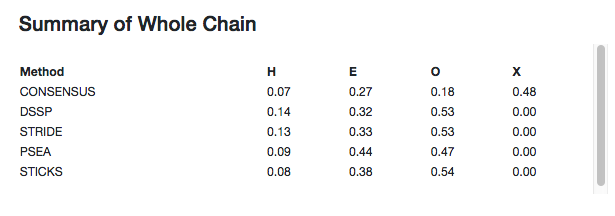
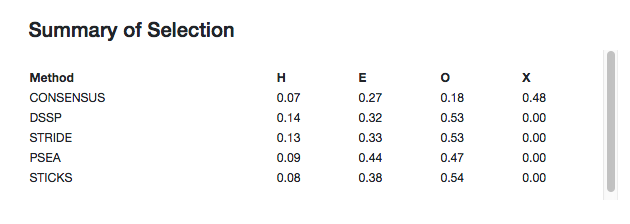
Summaries of the secondary structure fractions of the whole chain and the user defined selection are shown at the top of the page. This includes each of the active assignment methods as well as the consensus. "H" is helix, "E" is strand, "O" is other and "X" is undefined (when no consensus could be obtained).
-
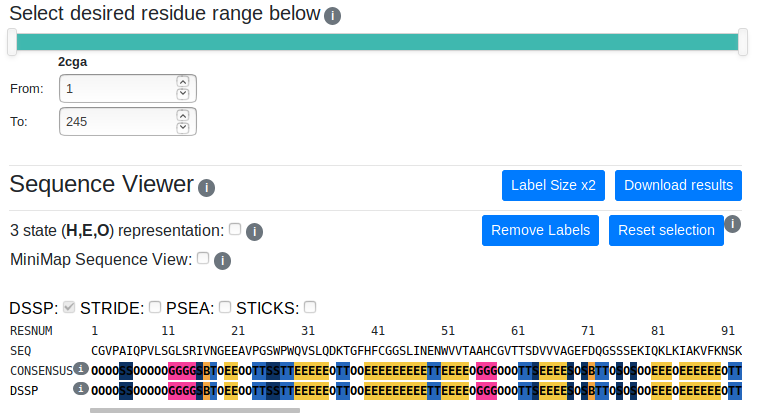
You can select the residue range to displace/analyse using the two-handed slider or the input boxes for greater precision. Deselected regions will be greyed out in the sequence viewer and will be hidden in the 3D viewer.

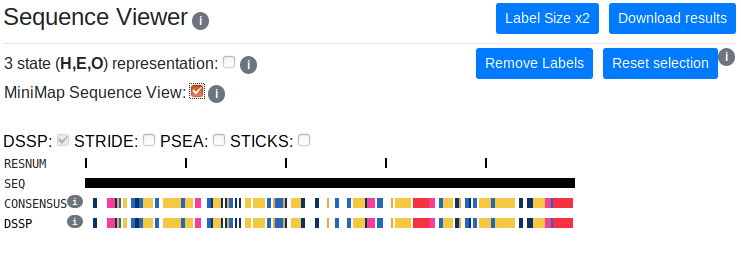
Sequence Viewer
-
The sequence viewer displays the residue number, the one-letter amino acid code and the consensus and individual method secondary structure assignments. Long sequences can be viewed by scrolling horizontally.
-
Selection and deselection of specific residues can be achieved by clicking the residue of interest in the "SEQ" row. Multiple residues can be toggled by clicking and dragging along the sequence row, which will highlight affected residues in green.
-
Clicking a residue in any of the secondary structure rows will zoom and focus on that residue in the 3D viewer. Mousing over a residue will pop up a tooltip displaying the residue number.
-
The secondary structure rows are coloured according to each residues assigned class. For each method the class-colour mapping key can be seen by mousing over the i buttons.
-
You can select which methods to include in the analysis using the checkboxes. This will trigger recalculation of the consensus.
-
If an assignment method fails, the checkbox will be greyed out/disabled, and the method name will be coloured red.
-
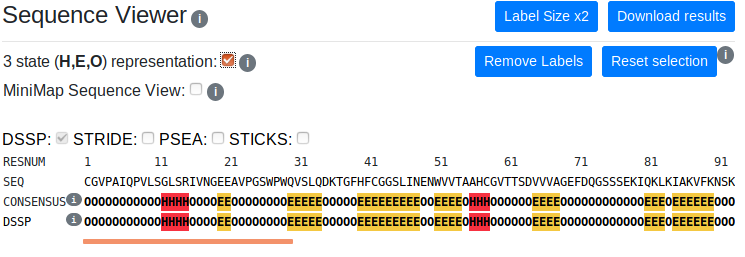
You can switch between full and three state representations of secondary structure using the checkbox. This will update the sequence viewer and 3D viewer colour schemes and trigger recalculation of the consensus. Mapping of full representation classes to the three state represenations for each method can be seen by mousing over the i buttons.
-
The MiniMap sequence view is useful if you are viewing large chains. It is a zoomed out view of the sequence and secondary structure assignments. All functionality of the full viewer is available in this mode.
-
Clicking the "Reset Selection" button will remove any selections applied by the user and recenter the protein in the 3D viewer.
-
Clicking the "Label Size" button will change the size of the labels in the 3D viewer.
-
Clicking the "Remove Labels" button will remove any labels added by the user in the 3D viewer.
-
An ASCII .csv file containing the results of the analysis can be downloaded by clicking the "Download Results" button. The files contents will reflect the choice of methods and selections made by the user.
3D Viewer
-
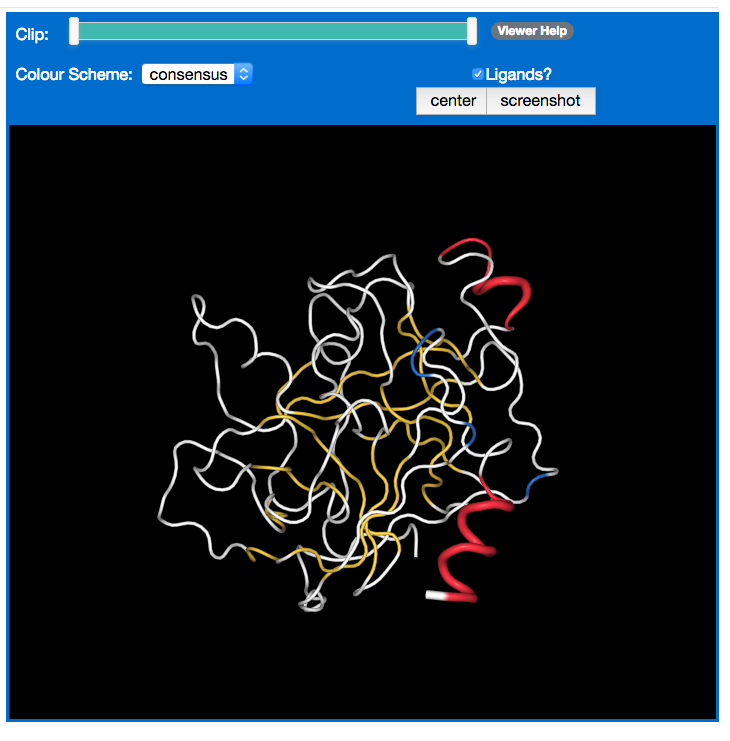
The 3D model of the currently selected chain is displayed in the 3D viewer.
-
The molecule can be rotated by clicking and dragging with the left mouse.
-
Holding the right mouse and moving the cursor pans the view.
-
Zooming can be achieved using the scroll wheel or equivalent e.g. two finger scrolling on a touchpad.
-
Mousing over an atom/residue will bring up a tooltip displaying information about the atom, residue and chain.
-
Holding Left-Ctrl and left clicking on an atom will label it with the residue name and number. Labels can be removed by the same method.
-
The clipping range of the camera can be controlled by using the two-handed slider.
-
The structure can be coloured according to its per residue secondary structure class from the consensus or the individual assignment methods. The colours are the same as that used in the sequence viewer. If a method has failed, it will not be available for colouring.
-
Display of ligands can be switched on and off using the checkbox.
-
The view can be centered on the whole molecule by clicking the "Center" button, or on specific residues by clicking them with the middle mouse button/wheel.
-
A .png image of the current view can be downloaded by clicking the "Screenshot" button. The image will not have a black background - instead it will have a transparent background making it suitable for use as a figure in presentations or for publication.
Submitting a job
Input: Two PDB files uploaded from your computer or two PDB 4-letter codes, or one PDB file (uploaded using the first file upload input) and one PDB code (entered into the first input), then click Submit. The Submit button will be disabled unless a viable combination of the above is input/uplaoded.
Pairing chains for analysis
-
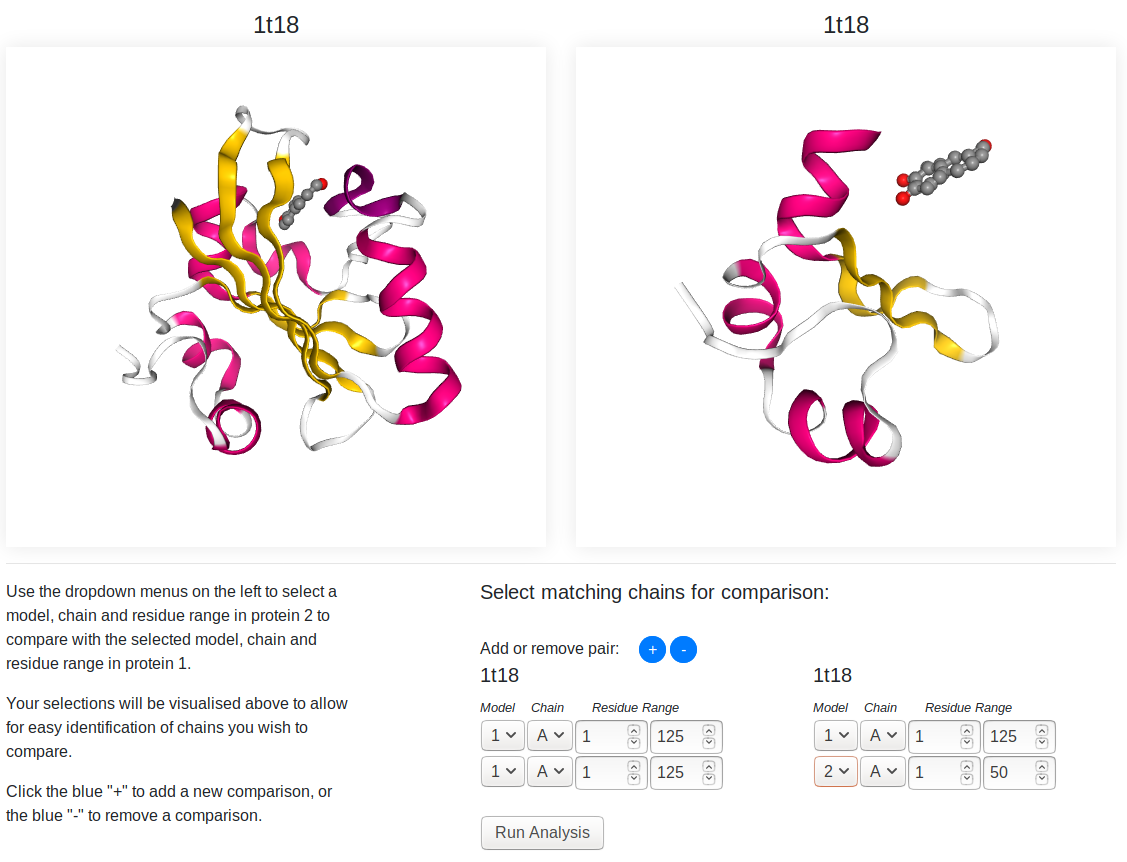
This page is used to choose the pairs of chains you want to analyse.
-
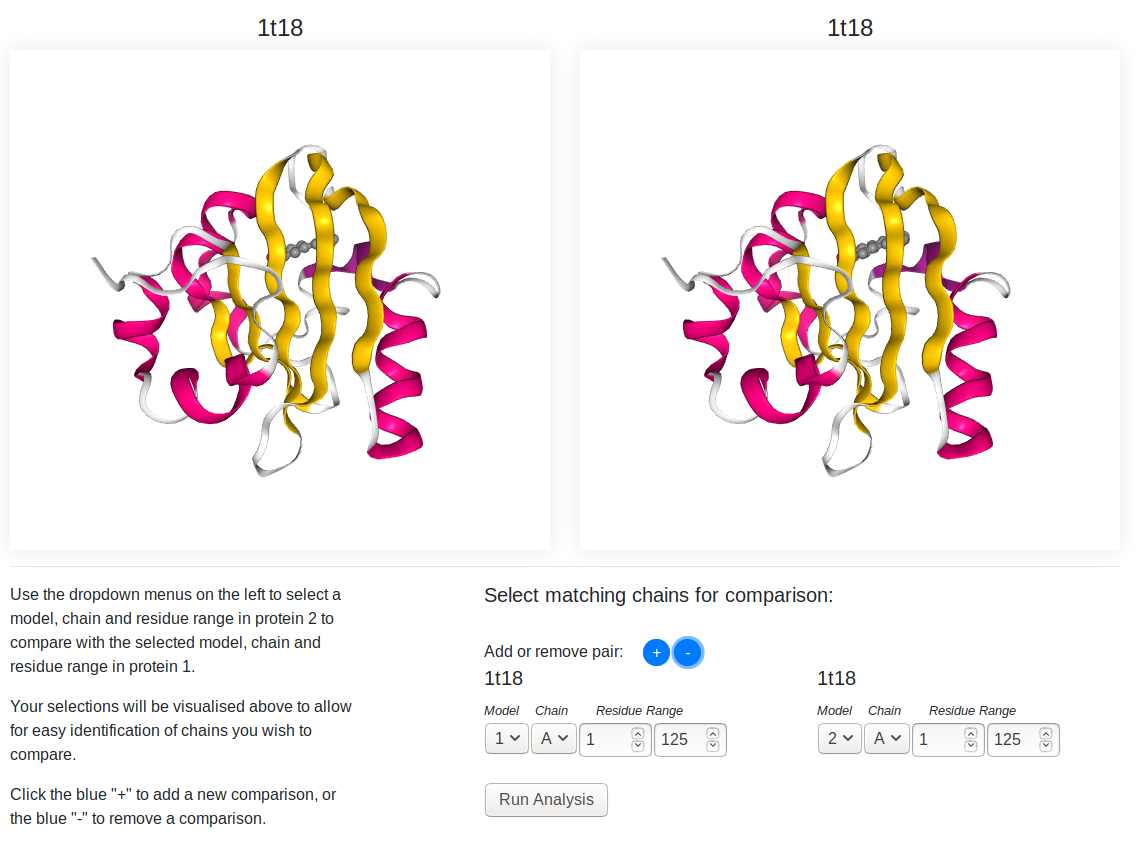
The two structures submitted for analysis are shown in the 3D viewers, protein A (the first protein submitted) on the left, protein B on the right. You can rotate (left click and drag), pan (right click and drag) and zoom (mouse wheel/two finger scroll) the structures using your mouse or touchpad.
-
On the bottom right, you can select the pairs you want to analyse. You can choose the model, chain and residue range of each protein you wish to analyse. If you want to do additional comparison, you can click the blue "+" to add another row to the form. Clicking the blue "-" will remove the last comparison added from the list.
-
Changing the selection for either protein will alter the 3D representations of the proteins. Only the selected model/chain/residue range (and any associated ligands) will be visible in the preview windows, and the view will automatically center to this selection. This provides an easier view for checking the folds of these chains and ensuring they are the ones you wish to compare.
-
Once you have chosen your pairs, click the "Run Analysis" button to be taken to the results page.

Results Page
-
Choose which pair you wish to see the results for using the dropdown menu on the top-left.
-
Below this you can choose to include or remove missing residues from the analysis. These are automatically parsed from the PDB header if present and are classified as "other".
-
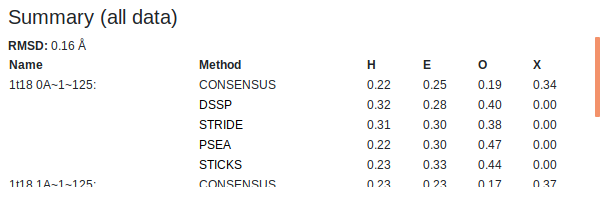
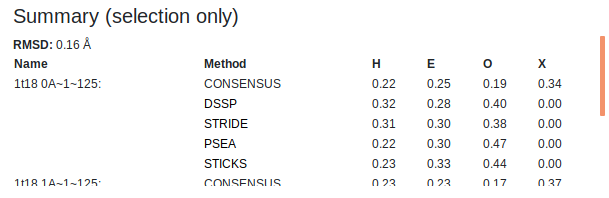
Summaries of the secondary structure fractions of the whole chain and the user defined selection for both analysed chains are shown at the top of the page. This includes each of the active assignment methods as well as the consensus. "H" is helix, "E" is strand, "O" is other and "X" is undefined (when no consensus could be obtained). Above these tables the RMSD (in Angstroms) of the aligned chain backbones is shown for the whole chains as well as the user defined selection.
-
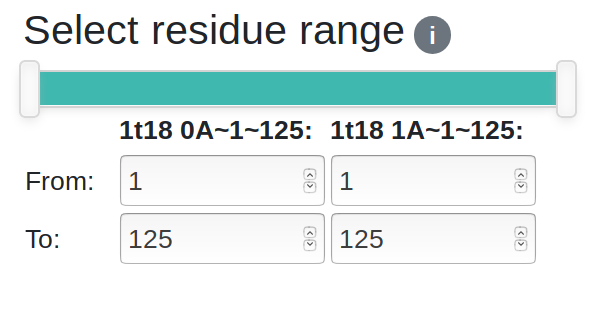
You can select the residue range to display/analyse using the two-handed slider or the input boxes for greater precision. There are two sets of input boxes, one for each chain. Changing any of the inputs will affect the selections of both chains. Deselected regions will be greyed out in the sequence viewer and will be hidden in the 3D viewer.
Sequence Viewer
-
The sequence viewer displays the residue number, the one-letter amino acid code and the consensus and individual method secondary structure assignments for both chains, aligned by sequence or by structure depending on which alignment had the least gaps. It also includes the differences found in secondary structure assignments between both chains, as well as the per-residue deviations of aligned C-alpha atoms. Long sequences can be viewed by scrolling horizontally. Gaps in the alignment are indicated by "-".
-
The type of alignment method chosen to generate the optimal comparison is indicated just below the range selection input. It will either be "Sequence" or "Structural".
-
Selection and deselection of specific residues can be achieved by clicking the residue of interest in the "SEQ" row. Multiple residues can be toggled by clicking and dragging along the sequence row, which will highlight affected residues in green. This will effect the selection in both chains.
-
The resolution of each structure is indicated in the same row as the B-factors. If the structure does not have a resolution (e.g. it is an NMR structure, or an uploaded file does not contain this information) it will display "N/A".
-
The B-factor rows are coloured according to each residues percentile rank compared to other structures with the same resolution's average B-factors. This can be used as an indication of how significant the analysis for a specific residue is. For example, if a residue is coloured red in the B-factor row, that indicates that residue's B-factor is in the 95th percentile or higher for structures with the same resolution. This tells you that the atom positions for that specific residue are poorly resolved compared to what might be expected, so you might decide that any differences found for this residue are not reliable. Conversely, if it was blue that indicates the B-factors are much lower than expected for a structure with that resolution, so a difference is much more likely to be significant. The colour scale can be seen by mousing over the i buttons. If the structure being analysed does not have B factors e.g. it is an NMR structure or a computationally generated model, the B-factor row will be coloured black and the "BFACTOR" row label will be coloured red.
-
Clicking a residue in any of the secondary structure rows or the B-factor row will zoom and focus on that residue in the 3D viewer. Mousing over a residue will pop up a tooltip displaying the residue number.
-
The secondary structure rows are coloured according to each residues assigned class. For each method the class-colour mapping key can be seen by mousing over the i buttons.
-
The difference row is coloured to indicate the proportion of currently selected methods that classified that residue's secondary structure differently. Black indicates there were no differences. A scale from yellow to red indicate differences were found - red indicating all methods predicted a different class. Gaps in the alignment are coloured black.
-
Selection and deselection of differences can be achieved by clicking the residue of interest in the "DIFFERENCES" row. Multiple residues can be toggled by clicking and dragging along the sequence row, which will highlight affected residues in green. This can be useful to remove differences that might not be of interest to you from the protein in the 3D viewer for production of figures or easier analysis. Deselecting differences in this manner is seperate to the main selection functionality - the residue will still be visible in the 3D viewer and only the specific residue in the difference row will be greyed out in the sequence viewer. However, the sidechain representation of that residue will not appear in the 3D viewer until you reselect it in the sequence viewer, or switch on visualisation of all sidechains using the "Sidechains?" checkbox. The residue will no longer be coloured as a difference in the 3D viewer if deselected, instead being coloured white.
-
The C alpha distances row is coloured to indicate the per-residue distance between aligned C-alphas. This scale goes from white (0 Angstrom distance) through yellow to red (>5 Angstrom distance). Gaps in the alignment are coloured black.
-
The side chain distances row is coloured to indicate the per-residue distance between aligned sidechains. This scale goes from white (0 Angstrom distance) through yellow to red (> (75% of the worst resolution structure being compared) Angstrom distance). Gaps in the alignment are coloured black. Only residues with the same amino acid type are compared in this manner, else the distance is set to 0.
-
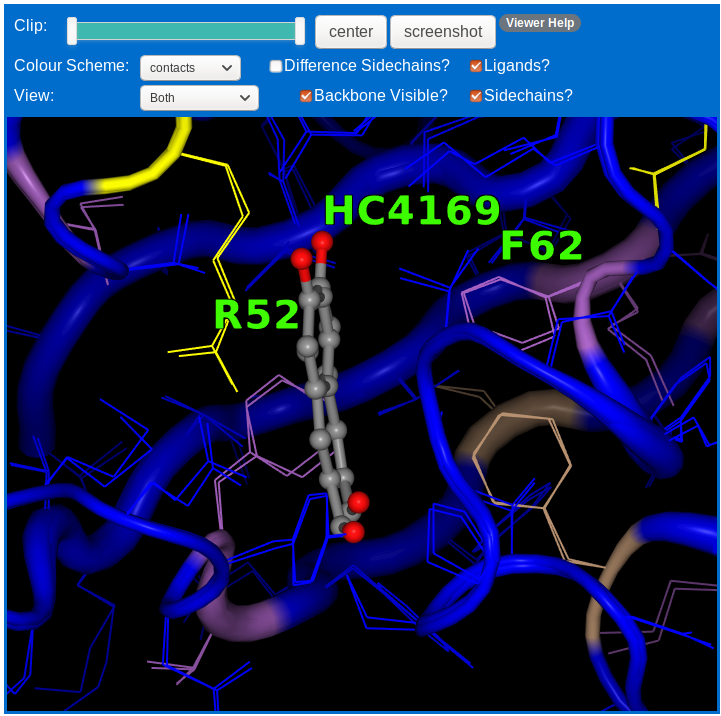
The contacts row is coloured to indicate the change in contacts between aligned sidechains. This is the number of contacts not found in BOTH the chains. This scale goes from blue (no change in contacts) through yellow to red (> 6 different contacts). Gaps in the alignment are coloured black.
-
You can select which methods to include in the analysis using the checkboxes. This will trigger recalculation of the consensus and differences.
-
If an assignment method fails, the checkbox will be greyed out/disabled, and the method name will be coloured red.
-
You can switch between full and three state representations of secondary structure using the checkbox. This will update the sequence viewer and 3D viewer colour schemes and trigger recalculation of the consensus and differences. Mapping of full representation classes to the three state represenations for each method can be seen by mousing over the i buttons.
-
The MiniMap sequence view is useful if you are viewing large chains. It is a zoomed out view of the sequence and secondary structure assignments. All functionality of the full viewer is available in this mode.
-
Clicking the "Remove Labels" button will remove any labels added by the user in the 3D viewer.
-
Clicking the "Label Size" button will change the size of the labels in the 3D viewer.
-
Clicking the "Reset Selection" button will remove any selections applied by the user (including difference deselections) and recenter the protein in the 3D viewer.
-
Clicking the "Invert diff. Sel" button will make all differences currently deselected selected, and vice versa.
-
Clicking the "Toggle all diff. ON/OFF" button will turn all the differences on or off. The button text will change to indicate whether a click will turn the differences on or off i.e. will say "Toggle all diff. ON" if clicking will turn all the differences on.
-
An ASCII .csv file containing the results of the analysis can be downloaded by clicking the "Download Results" button. The files contents will reflect the choice of methods and selections made by the user. Click here for an example.



3D Viewer
-
The 3D model of the currently selected, aligned pair of chains is displayed in the 3D viewer.
-
The molecules can be rotated by clicking and dragging with the left mouse.
-
Holding the right mouse and moving the cursor pans the view.
-
Zooming can be achieved using the scroll wheel or equivalent e.g. two finger scrolling on a touchpad.
-
Mousing over an atom/residue will bring up a tooltip displaying information about the atom, residue and chain.
-
Holding Left-Ctrl and left clicking on an atom will label it with the residue name and number. Labels can be removed by the same method.
-
The clipping range of the camera can be controlled by using the two-handed slider.
-
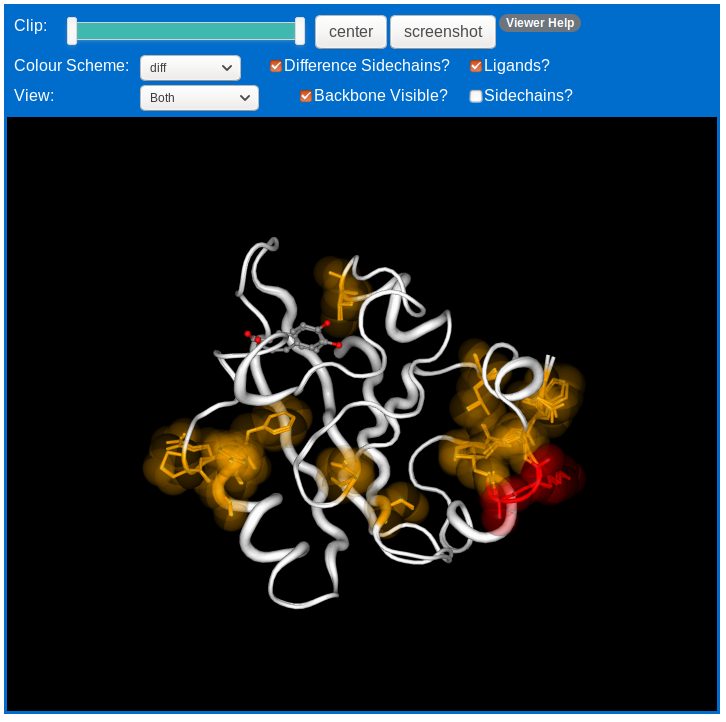
The structure can be coloured to show a variety of information. It can be coloured by the differences found, or by the per-residue C-alpha or sidechain distances, or by difference in contacts. It can be coloured by its per residue secondary structure class from the consensus or the individual assignment methods. It can also be coloured by chain (protein A chain in green, B chain in purple), B-factor percentile rank or by the CPK colour scheme to color displayed sidechains by atom-type. The colours are the same as that used in the sequence viewer. The default colour-scheme is the difference scheme.
-
By default the sidechains of residues for which a difference has been found are displayed as licorice representation with a translucent spacefill. You can toggle this on and off using the "Difference Sidechains?" checkbox.
-
You can also switch on and off the display of ALL sidechains using the "Sidechains?" checkbox. This and the "Difference Sidechains?" checkboxes work independently - so checking "Difference Sidechains?" and unchecking "Sidechains?" will result in the sidechains for residues where a difference has been found being displayed.
-
If you only wish to display the residues that are different and remove the rest of the protein backbone, you can do so using the "Backbone Visible?" checkbox.
-
Using the "View" dropdown menu, you can choose to view both, or either of the chains being compared alone. Chains are identified by the file name or PDB code of the protein they are from.
-
Display of ligands can be switched on and off using the checkbox.
-
The view can be centered on the whole molecule by clicking the "Center" button, or on specific residues by clicking them with the middle mouse button/wheel.
-
A .png image of the current view can be downloaded by clicking the "Screenshot" button. The image will not have a black background - instead it will have a transparent background making it suitable for use as a figure in presentations or for publication.
Q. How is the difference between protein structures calculated?
A. The difference calculated by 2StrucCompare is represented behind the scenes by a value between 0 and 1 for each residue indicating the proportion of secondary structure assignment methods that agree on the classification of said residue for both protein chains in the alignment. This value is recalculated dynamically when you add or remove the results from the individual methods to and from the analysis using the checkboxes on the results page. For example, if you had just DSSP active (which is the default) and the nth residue in the alignment of protein A had an assignment of "H" and the equivalent aligned residue in protein B had an assignment of "G", the difference at this residue would be 1.0 - the maximum possible difference value, as all methods disagree. If you then added STRIDE to the analysis, and the stride assignments for A and B were "H" and "H", the STRIDE assignments agree - this means half the methods have a disagreement and the difference value at this position in the alignment would be (1+0)/2 = 0.5. This is reflected in the colour scale shown in the sequence viewer, and the raw difference values between 0 and 1 are available in the ASCII text file of results downloadble from the results page.
Q. How is the consensus calculated?
A. The consensus is calculated for 2Struc and 2StrucCompare in the same manner - for each chain active on the results page (for 2Struc, one chain, for 2StrucCompare, two chains) a consensus is produced. For each residue in the chain, the secondary structure assignments for that residue are collected and a determination is made if 50% or more of the classes are the same. If there is a clear majority for a particular class, that assignment becomes the consensus assignment for that residue. If there is no majority, the assignment is considered to be undefined for the consensus sequence, indicated by an "X".
Q. How is the C-alpha distance calculated?
A. The per-residue C-alpha distance is calculated between aligned residues by finding the euclidean distance between the coordinates of each residues C-alpha atoms.
Q. How is the sidechain distances calculated?
A. For aligned residue pair, r1 and r2, if the amino acid type of both residues was the same, the deviation of sidechains was calculated. The coordinates of all atoms in the sidechain (excluding backbone atoms) are collected. To correct for large deviations in backbone position the vector, v, required to translate the C alpha atom coordinates of r1 (x1, y1, z1) to the C alpha atom coordinates (x2, y2, z2) of r2 is calculated. The atomic coordinates of r1 are then translated using v, before being used in the subsequent side chain deviation calculation. The Euclidean distance between equivalent atoms in r1 (corrected using v) and r2 are calculated as above (See C alpha deviations). The maximum interatom distance calculated (in Angstroms) is taken as the sidechain deviation for the aligned pair r1 and r2.
Q. How are the contacts calculated?
A. For the aligned residue pair r1 and r2, the contacts each make within their parent chains were calculated. The NeighbourSearch function in the Bio.PDB python library was used (Hamelryck & Manderick (2003), Cock et al. (2009)). This function finds all residues within a radius of a given query position using K-D trees. A radius of 4 Angstroms was used for 2StrucCompare, and the coordinates of the all atoms in r1 or r2 were used as the query position to calculate the sets of contacts c1 and c2 respectively. Once the contact sets were calculated, the difference in contacts made (i.e. contacts not found in both c1 and c2) was found by counting the number of contacts left after taking the union of c1 and c2 and subtracting the intersect of c1 and c2.
Q. How do we choose which alignment method to use?
A. We envisioned 2StrucCompare would be used for two different kinds of analysis - comparing two structures with identical or near-identical sequences under different conditions (e.g. change in pH, binding partner, a point mutation) or for comparing analagous structures which may have very similar folds but very low sequence identity. To that end, we perform both a sequence alignment using the Needleman-Wunsch global alignment algorithm and a structural alignment using TMAlign. The two alignments that are produced are then scored based on the number of gaps that are opened (+1 point), and the number of extensions to those opened gaps (+0.1 point). The structure with the LOWEST score (usually the alignment with the least number of gaps) is then chosen. If the sequence alignment is chosen, that is what is displayed in the Sequence Viewer, and the sequence alignment is used as input to TMAlign to produce the structural alignment you can see in the 3D viewer. If the structural alignment wins, that will be displayed in both the Sequence Viewer and the 3D viewer. The type of alignment method chosen is also indicated on the results page, in the Sequence Viewer pane.
Q. How should I intepret the BFACTOR row?
A. The BFACTOR row is there to provide you with a metric to judge how significant the calculated differences are. A residue with a low average B-factor suggests we are confident in its atomic positions, therefore we can be confident in the results that rely on this factor e.g. calculation of hydrogen bonds during secondary structure assignment. Conversely, a high B-factor would reduce our confidence in the result. We chose to display B-factors by their percentile rank compared to all other structures in the PDB with the same resolution as your input. This provides an indication of whether the residue you are interested in has a good or bad B-factor compared to structures of the same quality. For example, if you see a residue is coloured red in the BFACTOR row, that suggests it is in the 95th or higher percentile - it has a higher or equal b-factor as 95% of the residues in structures with the same resolution. If it was coloured green, that indicates its in approximately the 50th percentile, which means its B-factor is average for that resolution, and if it is dark blue, that indicates it is in a lower percentile rank (e.g. 10th) which means it has very low B-factors compared to other strucutures at that resolution. Some types of proteins tend to have higher B-factors than others due to the environments they are solved in, or the way B-factors are calculated for them (e.g. Membrane proteins) - in all cases, the BFACTOR row only exists as a guide to significance and you should use your discretion when deciding what results you have confidence in.
Q. Should I choose 3 state or full representation?
A. The consensus and differences calculated can change significantly when changing representation. The full representation, due to its finer partitioning of secondary structure into a greater number of classes (although PSEA only defines 3 classes in the full representation) allows for the detection of more subtle changes in structure - e.g. a change from alpha helix to 3/10 helix may not look like a large change to the naked eye, but indicates a real alteration in side chain position and hydrogen bonding patterns has occurred which could be biologically relevant. However, the full representation might be over-sensitive in some scenarios, or the user might not be as interested in differences of turn to other, for example. In that case, the 3 state representation reduces the complexity of the analysis while still preserving the large changes e.g. helix to strand or other that would have the biggest effect.
Q. I submitted my PDB files but I got an error. Why did that happen?
A. There are several reasons the analysis might not have worked, usually due to problems with the format or content of the input files. Below is a short list of the most common:
- The code you entered does not exist - check on the RCSB PDB website that the code you entered is correct and the entry has data associated with it.
- The PDB file or code provided does not contain protein - if there is no protein in the file, there is nothing to analyse. Check that the file contains at least one chain that is protein, and that the code you entered is correct.
- The PDB file or code provided has formatting issues. Check the wwPDB website for a guide to the format.
- You provided two identical files to 2StrucCompare - this could mean you provided the same file twice, or you provided different files which have identical atomic coordinates. The last case is rare, but we did come across it in testing with files from the PDB.
Q. An error message said "Incorrect extension for file: XXXX.XXX. Only *.pdb, *.ent and *.cif are allowed.". What do I do?
A. The server only accepts PDB format files with .pdb or .ent file extensions, or MMCIF formatted files with a .cif extension. If you have, for example, a .txt file that contains a correctly formatted PDB structure, you will need to change the file extension from .txt to .pdb or .ent. Below are a number of ways you can do this depending on your operating system:
- Mac OSX: Find the file in Finder, right click the file to bring up the context menu, select "Get Info" and change the extension in the "Name & Extension" box in the dialog that pops up.
- Linux (Ubuntu/Debian): Find the file in your distributions file explorer. Right click to bring up the context menu, click "Rename.." and change the extension. Other distributions of Linux may have different methods to accomplish this.
- Windows 10: Open File Explorer and locate the file. If you cannot see the extension in the file name, you may need to enable the display of file extensions. Click the "View" ribbon and make sure the checkbox for "File name extensions" is checked. Now click "Options" and select "Change folder and search options" from the dropdown menu. Make sure the option "Hide extensions for known file types" is unchecked/unticked. Once you are able to see the extension in the file name in the File Explorer window, you can right click on the file, choose "Rename" from the context menu and change the file extension.
Ensure that you choose the right extension out of .pdb, .ent or .cif to match the format of the structural information stored in your file, otherwise the analysis will fail.
Q. My structure has modified/non-standard amino acids, is this alright?
A. Some of the programs that we run to perform the analysis are old and do not handle modified residues or in-polymer HETATM entries very well. Comparing structures that contain non-standard amino acids that are part of the protein chain (i.e. not free ligands) will work, however you may find these specific residues are absent from the analysis shown in the Sequence Viewer as we were not able to obtain results for them from all the required programs. This will not adversely affect the accuracy of the results for the displayed residues, however. If you wish to modify your PDB files to alleviate this issue, (if you believe it will not adversely affect the accuracy of the results) you could change the non-standard residue to its "parent" residue i.e. MSE to MET, and change the line identifier in your file from HETATM to ATOM. This should then allow results to be obtained for this position. If you have continued issues with analysing a structure with a modified residue, please don't hesitate to email us with the PDB code or file, and the identity of the residue.
Q. How long will my results be available for on the site?
A. The results page will be available to view for 48 hours from time of submission. After that period, all user data and analysis is deleted from our servers and you will have to resubmit the structures to see the results again.
Q. Can I use this website on my phone/tablet/touchscreen device?
A. The site was designed primarily to be used with a mouse or mouse-like pointer device. You can run the analysis using a touchscreen device like a phone or tablet and view your results just like on a desktop or laptop, but you may run into issues with some of the functionalities of the 3D Viewer and Sequence Viewer.
Q. I have a suggestion for the site. How can I contact you?
A. If you have a suggestion for the site - the analyses we run, an idea for a new kind of difference metric you would like to be able to perform using our service, user experience - we would welcome your suggestions as we are interested in actively improving all the services we provide. Please contact the developer Dr Elliot Drew with any thoughts or suggestions.
